|
Peak calling was done with the clc genomics workbench For each sample the appropriate input control was considered. Peak calling using a P-value threshold of 0.05. Mapping of reads to the Arabidopsis genome (TAIR10) using the clc genomics workbench and default settings. 10 µL of each indexed library were pooled and the final library was purified using AMPure resin.įiltering of uniquely mappable reads using the clc genomics workbench (version 8.5.1, Quiagen) 1 ng of each sample and input was used for library preparation, 11 amplification cycles were applied to amplify the libraries. 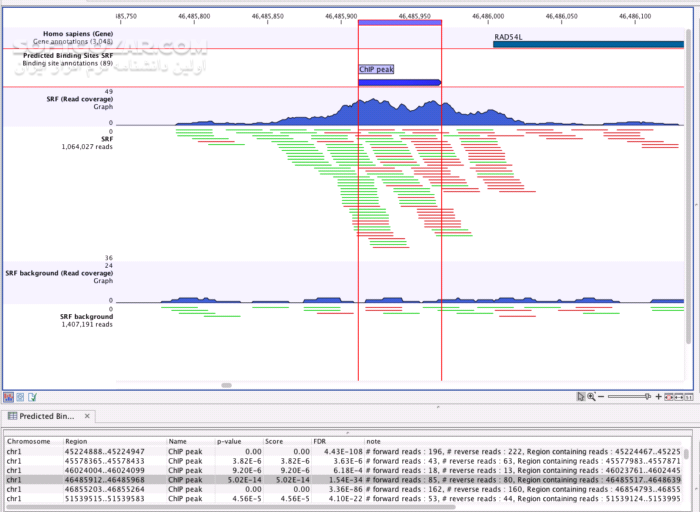 
Libraries were prepared using the MicroPlex Library Preparation kit v2 (Diagenode, C05010014) according to the manual. GEO help: Mouse over screen elements for information.Ĭhip antibody: anti-H3K9/14ac (Diagenode, C15410200)Ģ50 µM GSNO, 250 µM GSH, 250 µM GSNO / 500 µM cPTIO, 5 µM TSA for 3h and 16hĪpproximately 20 µL of seeds (dry volume) were grown in sterile 1x MS supplemented with 1% sucrose under moderate shaking (120 rpm) and short-day conditions (10h light/ 14h dark, light intensity 130 µmol s-1 m-2)Ĭhromatin was isolated, sheared and subjected to immunoprecipitation using the Plant ChIP-seq Kit (Diagenode, C01010150) according to the manual.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
